November 19, 2023
While next generation sequencing (NGS) is receiving all the headlines, traditional Sanger sequencing is carrying on the grunt of the work in day-to-day laboratory analysis. Here at Agri-Analysis we routinely use DNA sequencing, combined with other techniques such as PCR or RT-PCR, to help answer growers questions in number of areas:
- Pathogenic strains of Agrobacterium vitis are known to be the causal agent for crown diseases. PCR alone is insufficient for the reliable identification of A. Vitis. We use a combination of PCR followed by DNA sequencing for reliable identification of A. vitis in both vineyards and nursery materials.
- Confirmation of fungicide resistance of powdery mildew to Qoi (Franc 11 ) and DMI (frac 3) fungicides in grapevines has been increasingly requested by growers when they experience field failures when using these fungicides. PCR followed by DNA Sequencing can help us pin-point the exact mutations in PM genes attributable to resistance.
- Grapevine leafroll associated virus type 3 (GLRaV-3) is the most devastating virus in grape production worldwide. Because GLRaV-3 genomes are highly variant, some strains can escape detection. We have used PcR followed by DNA sequencing for early discovery and testing of the emergence of a new grapevine leafroll associated virus type 3 (GLRaV-3) variant in commercial vineyards. Our work preceded the discovery of the same by NGS by several years. This provided us with a competitive advantage by being able to test for the new variant several years before its identification and publication by researchers.
- Cherry X disease, attributable to candidatus phytoplasma, is an economically impactful disease in the cherry producing regions of the U.S. We have used nested-PCR followed by DNA sequencing for identification and confirmation of phytoplasma infection in cherry orchards in California.
- We have used PCR followed by DNA sequencing for the identification and confirmation of infection by a number of known viruses in Apple and Cherry orchards such as Apple Stem Grooving virus, Apple chlorotic leaf spot virus, prunus dwarf virus and prunus necrotic ringspot virus.
- We have used PCR followed by DNA sequencing identification and confirmation of hop latent viroid (HLVd) which is the biggest concern for cannabis and hop growers worldwide although most HLVd-infected plants can remain asymptomatic.
- We have used PCR followed by DNA sequencing identification and confirmation of new infection of a known virus such as Pinot Gris virus (GPGV) for the first time in California commercial vineyards;
- We have used PCR followed by DNA sequencing for clade typing and phylogenetic analysis of viruses in infected plants. For example, we found that GPGV infections in California vineyards, to date, are attributable to the non-symptomatic clade of GPGV which is less economically damaging than its symptomatic variant.
- Sometimes, we are asked to conduct clade typing analysis of grapevine red blotch virus in infected vines to aid growers tracking where the infection may have initiated and spread. There are two known clades of GRBaV which vary by about 7% in genetic sequences but otherwise are phenotypically identical.
Please email me at apwei@agri-analysis.com with any questions and comments on this and other topics.
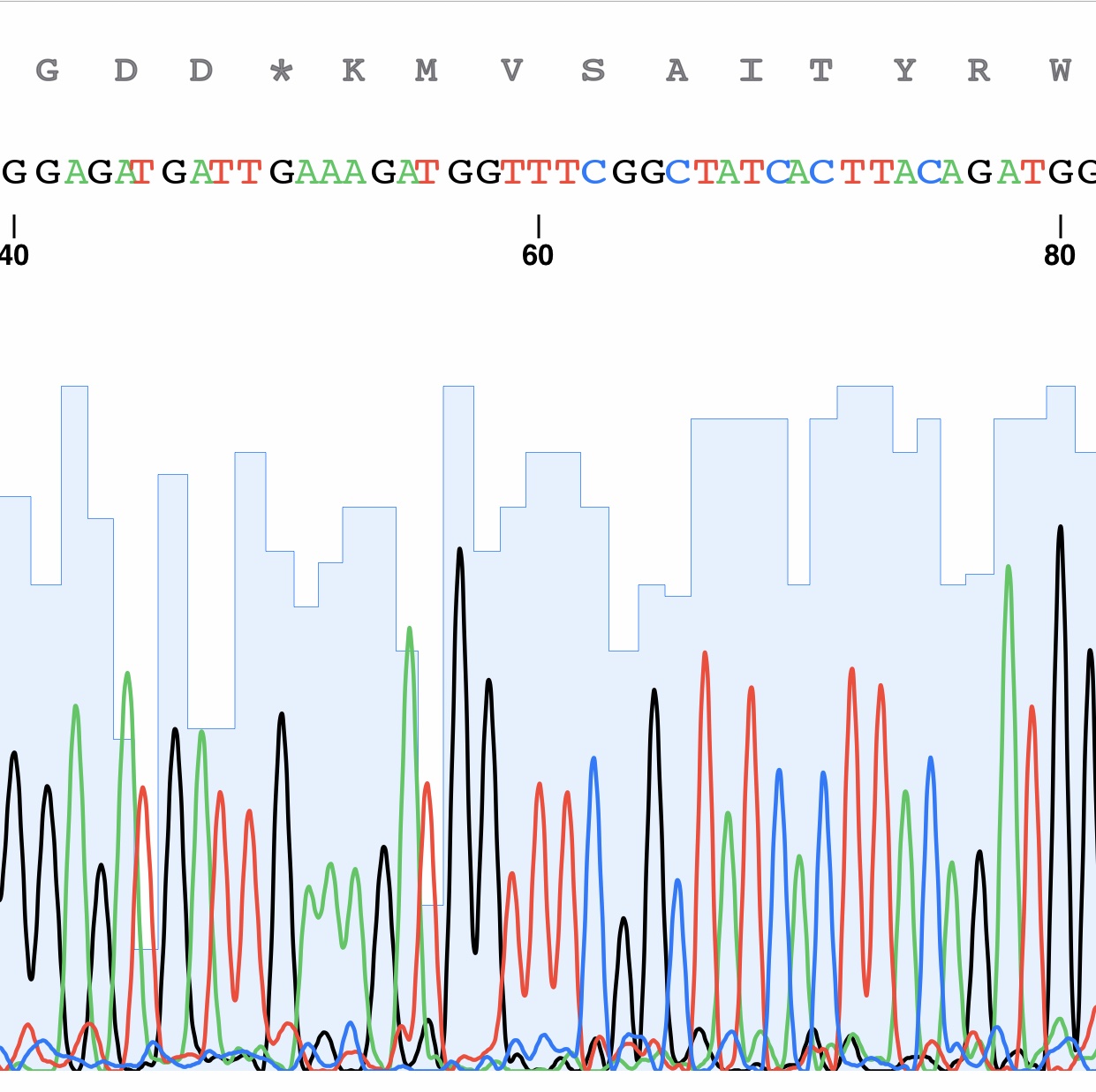
An example of a DNA sequencing chromatogram (partial).
00

